AngularAutoCorrelationsTwoAxes v1#
Summary#
Calculates the angular auto-correlations of molecules in a simulation along two user-defined axes. The first axis is defined by the vector connecting the average position of species two and the average position of species one (user input). The second axis is perpendicular to axis 1 and is constructed by considering one arbitrary atom of species 3 (user input). Timestep must be specified in femtoseconds.
Properties#
Name |
Direction |
Type |
Default |
Description |
|---|---|---|---|---|
InputFile |
Input |
string |
Mandatory |
Input .nc file with an MMTK trajectory |
Timestep |
Input |
string |
1.0 |
Time step between two coordinates in the trajectory in femtoseconds |
SpeciesOne |
Input |
string |
Specify the first species, e.g. H, He, Li… |
|
SpeciesTwo |
Input |
string |
Specify the second species, e.g. H, He, Li… |
|
SpeciesThree |
Input |
string |
Specify the third species, e.g. H, He, Li… |
|
OutputWorkspace |
Output |
Mandatory |
Output workspace name |
|
OutputWorkspaceFT |
Output |
FT |
FT Output workspace name |
Description#
Loads a netcdf file generated by nMoldyn containing MMTK format trajectories. The algorithm calculates angular auto-correlations of molecule in the simulation along a user-defined axis. The trajectory file must therefore contain molecule definitions. The first axis vector is drawn from the average position of atoms of type SpeciesOne to the average position of atoms of type SpeciesTwo. The second axis vector is drawn by extracting the component, orthogonal to the first axis, of the vector connecting the average position of atoms of type SpeciesTwo and an arbitrary atom of type SpeciesThree.
Example#
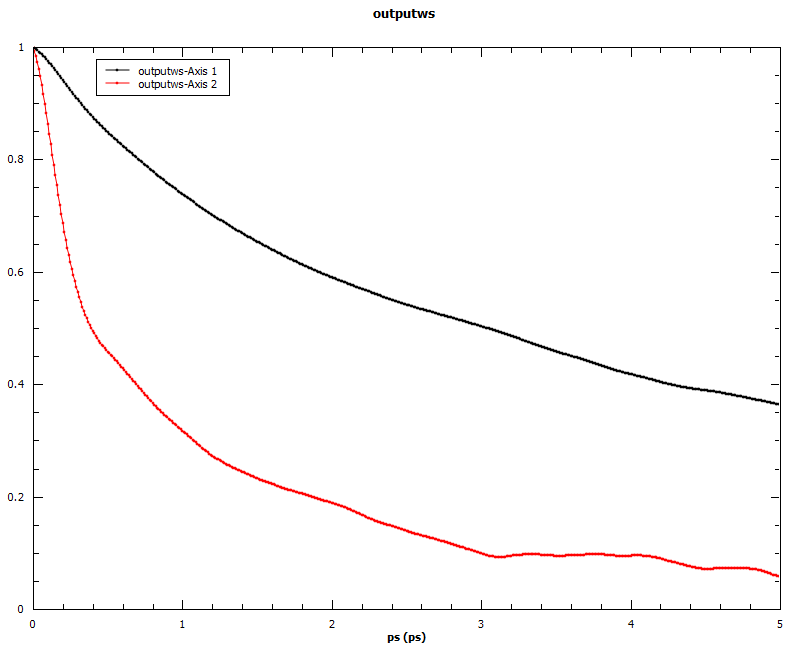
Angular auto-correlations calculated for methyliodide.
Usage#
output_ws, output_ws_ft = AngularCorrelationsTwoAxes(InputFile = 'trajectory_methyliodide.nc',
Timestep = '10.0',
SpeciesOne = 'C',
SpeciesTwo = 'I',
SpeciesThree = 'H')
Categories: AlgorithmIndex | Simulation

