\(\renewcommand\AA{\unicode{x212B}}\)
SaveNXSPE v1¶
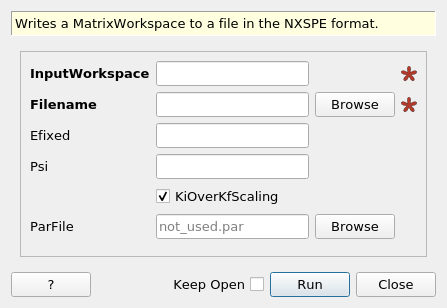
SaveNXSPE dialog.¶
Summary¶
Writes a MatrixWorkspace to a file in the NXSPE format.
See Also¶
Properties¶
Name |
Direction |
Type |
Default |
Description |
|---|---|---|---|---|
InputWorkspace |
Input |
Mandatory |
The name of the workspace to save. |
|
Filename |
Input |
string |
Mandatory |
The name of the NXSPE file to write, as a full or relative path. Allowed extensions: [‘.nxspe’] |
Efixed |
Input |
number |
Optional |
Value of the fixed energy to write into NXSPE file. |
Psi |
Input |
number |
Optional |
Value of PSI to write into NXSPE file. |
KiOverKfScaling |
Input |
boolean |
True |
Flags in the file whether Ki/Kf scaling has been done or not. |
ParFile |
Input |
string |
not_used.par |
If provided, will replace detectors parameters in resulting nxspe file with the values taken from the file. Should be used only if the parameters, calculated by the [[FindDetectorsPar]] algorithm are not suitable for some reason. See [[FindDetectorsPar]] description for the details. Allowed extensions: [‘.par’, ‘.phx’] |
Description¶
Saves the given Matrix Workspace to a file in the NeXus-based ‘NXSPE’ format.
Restrictions on the Input Workspace¶
The input workspace must have units of Momentum Transfer (‘DeltaE’) and contain histogram data with common binning on all spectra.
Child Algorithms Used¶
The FindDetectorsPar v1 algorithm is used to calculate detectors parameters from the instrument description.
Usage¶
Example - Save/Load “Roundtrip”
import os
import numpy
# Create dummy workspace.
out_ws = CreateSimulationWorkspace(Instrument="IRIS", BinParams="0,500,2000")
out_ws.setY(0, numpy.array([10.0, 50.0, 30.0, 60.0]))
AddSampleLog(out_ws, 'Ei', LogText='321', LogType='Number', NumberType='Double')
out_ws.setDistribution(True)
file_path = os.path.join(config["defaultsave.directory"], "NXSPEData.nxspe")
# Do a "roundtrip" of the data.
SaveNXSPE(out_ws, file_path,Psi=32)
# By desigghn, SaveMXSPE does not store detector's ID-s. LoadNXSPE sets detector's ID-s to defaults.
# To compare loaded and saved workspaces here, one needs to set-up default detector's ID-s to the source workspace.
nSpec = out_ws.getNumberHistograms()
for i in range(0,nSpec):
sp=out_ws.getSpectrum(i);
sp.setDetectorID(i+1);
in_ws = LoadNXSPE(file_path)
ws_comparison_rez = CompareWorkspaces(out_ws,in_ws,1.e-9,CheckInstrument=False)
print("Contents of the first spectrum = {}.".format(in_ws.readY(0)))
print("Initial and loaded workspaces comparison is: {}".format(str(ws_comparison_rez[0])))
run = in_ws.getRun();
print("Loaded workspace has attached incident energy Ei={0:.1f} and rotation angle Psi= {1:.1f}deg".format(run.getLogData('Ei').value,run.getLogData('psi').value))
Output:
Contents of the first spectrum = [ 10. 50. 30. 60.].
Initial and loaded workspaces comparison is: True
Loaded workspace has attached incident energy Ei=321.0 and rotation angle Psi= 32.0deg
Note that LoadNXSPE v1 automatically applies the distribution flag to the loaded workspace. This is because all examples of workspaces saved to NXSPE format by the reduction algorithms are distributions (signal is count rate and should be multiplied by bin widths to get counts). SaveNXSPE does not require its input is a distribution, however, and the NXSPE format does not have a distribution flag.
Categories: AlgorithmIndex | DataHandling\Nexus | DataHandling\SPE | Inelastic\DataHandling
Source¶
C++ header: SaveNXSPE.h
C++ source: SaveNXSPE.cpp
