IMDWorkspace#
This is a Python binding to the C++ class Mantid::API::IMDWorkspace.
bases: mantid.api.Workspace, mantid.api.MDGeometry
The MD Workspace [MDWorkspace] (short for “Multi-Dimensional” Workspace) is a generic data structure holdings points (MDEvents) that are defined by their position in several dimensions.
See also
Description of MDWorkspace#
Dimensions: A MDWorkspace can have between 1 and 9 dimensions.
Each dimension is defined with a name, units, and minimum/maximum extents.
MDEvent: A MDEvent is simply a point in space defined by its coordinates, plus a signal (weight) and error.
The MDLeanEvent type contains only coordinates, signal and error.
The MDEvent type also contains a run index (for multiple runs summed into one workspace) and a detector ID, allowing for more information to be extracted.
The class is named MDEventWorkspace.
Structure#
The MDWorkspace is a container that can hold a large number of MDEvents. The events are organized into “boxes”: types are MDBox and MDGridBox. At the simplest level, an MDWorkspace will be a single MDBox with an unsorted bunch of events.
In order to allow for efficient searching and binning of these events, the boxes are organized into a recursive boxing structure (adaptive mesh refinement). During MDWorkspace construction, if a MDBox is found to contain too many events, it will be split into smaller boxes.
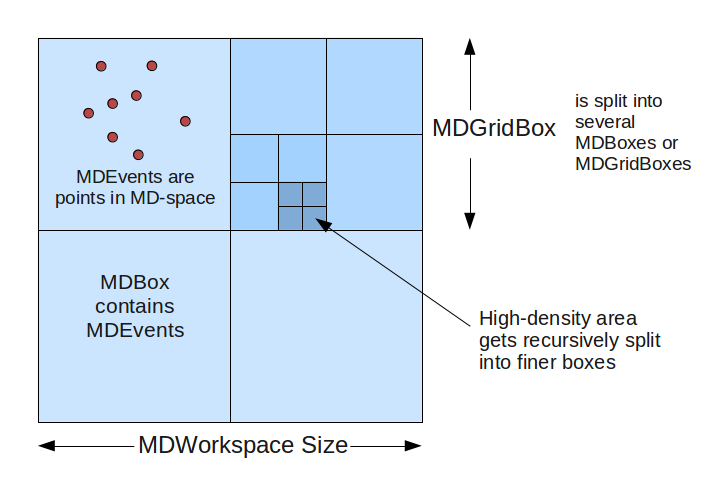
MDWorkspace_structure.png#
The threshold for splitting is defined in CreateMDWorkspace as the SplitThreshold parameter. Each parent box will get split into N sub-boxes in each dimension. For example, in a 2D workspace, you might split a parent box into 4x4 sub-boxes, creating 16 MDBoxes under the parent box (which becomes a MDGridBox). The level of splitting is defined in the SplitInto parameter.
Creating a MDWorkspace#
There are several algorithms that will create a MDWorkspace:
CreateMDWorkspace creates a blank MDWorkspace with any arbitrary set of dimensions.
CreateMD Creates an MDWorkspace in the Q3D, HKL frame.
ConvertToDiffractionMDWorkspace converts an
EventWorkspaceorWorkspace2Dfrom detector space to reciprocal space, for elastic single-crystal or powder diffraction experiments.ConvertToMD converts workspaces for inelastic experiments.
SliceMD takes a slice out of a MDWorkspace to create a new one.
LoadSQW converts from the SQW format.
File-Backed MDWorkspaces#
For workspaces with a large number of events that would not fit in memory, it is possible to use a NXS file back-end as a data store. The box structure will always remain in memory, but the underlying events will be stored in a file and retrieved only when required. This can be set at creation (CreateMDWorkspace) or when loading from a file, or an in-memory MDWorkspace can be converted to file-backed with the SaveMD algorithm.
Because of disk IO, file-backed MDWorkspaces are slower to process for some operations (e.g. binning or slicing). Some types of visualization and analysis, however, are just as fast with file-backed MDWorkspaces as their in-memory equivalent.
Viewing MDWorkspaces#
Right-click on a MDWorkspace and select:
Show Slice Viewer: to open the Sliceviewer, which shows 2D slices of the multiple-dimensional workspace.
Working with Table Workspaces in Python#
Accessing Workspaces#
The methods for getting a variable to an MDWorkspace is the same as shown in the Workspace help page.
If you want to check if a variable points to something that is an MDWorkspace Workspace you can use this:
from mantid.api import IMDEventWorkspace
mdws = CreateMDWorkspace(Dimensions=3, Extents='-10,10,-10,10,-10,10', Names='A,B,C', Units='U,U,U')
if isinstance(mdws, IMDEventWorkspace):
print(mdws.name() + " is a " + mdws.id())
Output:
mdws is a MDEventWorkspace<MDLeanEvent,3>
MD Workspace Properties#
For a full list of the available properties and operation look at the IMDEventWorkspace api page.
ws = CreateMDWorkspace(Dimensions='2', EventType='MDEvent', Extents='-10,10,-10,10',
Names='Q_lab_x,Q_lab_y', Units='A,B')
FakeMDEventData(ws, UniformParams="1000000")
print("Number of events = {}".format(ws.getNEvents()))
print("Number of dimensions = {}".format(ws.getNumDims()))
print("Normalization = {}".format(ws.displayNormalization()))
for i in range(ws.getNumDims()):
dimension = ws.getDimension(i)
print("\tDimension {0} Name: {1}".format(i,
dimension.name))
bc =ws.getBoxController()
print("Is the workspace using a file back end? {}".format(bc.isFileBacked()))
backEndFilename = bc.getFilename()
Dimensions#
As a generic multi dimensional container being able to access information about the dimensions is very important.
ws = CreateMDWorkspace(Dimensions='3', EventType='MDEvent', Extents='-10,10,-5,5,-1,1',
Names='Q_lab_x,Q_lab_y,Q_lab_z', Units='1\A,1\A,1\A')
FakeMDEventData(ws, UniformParams="1000000")
print("Number of dimensions = {}".format(ws.getNumDims()))
for i in range(ws.getNumDims()):
dimension = ws.getDimension(i)
print("\tDimension {0} Name: {1} id: {2} Range: {3}-{4} {5}".format(i,
dimension.getDimensionId(),
dimension.name,
dimension.getMinimum(),
dimension.getMaximum(),
dimension.getUnits()))
print("The dimension assigned to X = {}".format(ws.getXDimension().name))
print("The dimension assigned to Y = {}".format(ws.getYDimension().name))
try:
print("The dimension assigned to Z = {}".format(ws.getZDimension().name))
except RuntimeError:
# if the dimension does not exist you will get a RuntimeError
print("Workspace does not have a Z dimension")
# you can also get a dimension by it's id
dim = ws.getDimensionIndexById("Q_lab_x")
# or name
dim = ws.getDimensionIndexByName("Q_lab_x")
Accessing the Data#
To access the data of an MDWorkspace you need to convert it to a regular grid, or MDHistoWorkspace.
# Setup
mdWS = CreateMDWorkspace(Dimensions=4, Extents=[-1,1,-1,1,-1,1,-10,10], Names="H,K,L,E", Units="U,U,U,V")
FakeMDEventData(InputWorkspace=mdWS, PeakParams='500000,0,0,0,0,3')
# Create a histogrammed (binned) workspace with 100 bins in each of the H, K and L dimensions
histoWS = BinMD(InputWorkspace=mdWS, AlignedDim0='H,-1,1,100', AlignedDim1='K,-1,1,100', AlignedDim2='L,-1,1,100')
# Or you can also use CutMD, to define bin widths and the cut projection
from mantid.api import Projection
SetUB(Workspace=mdWS, a=1, b=1, c=1, alpha=90, beta=90, gamma=90)
SetSpecialCoordinates(InputWorkspace=mdWS, SpecialCoordinates='HKL')
projection = Projection([1,1,0], [-1,1,0])
proj_ws = projection.createWorkspace()
# Apply the cut with bin widths of 0.1 in H,K and L and integrating over -5 to +5 in E
out_md = CutMD(mdWS, Projection=proj_ws, PBins=([0.1], [0.1], [0.1], [-5,5]), NoPix=True)
Reference#
- class mantid.api.IMDWorkspace#
- clearOriginalWorkspaces((MDGeometry)self) None :#
Clear any attached original workspaces
- clone(InputWorkspace)#
Copies an existing workspace into a new one.
Property descriptions:
InputWorkspace(Input:req) Workspace Name of the input workspace. Must be a MatrixWorkspace (2D or EventWorkspace), a PeaksWorkspace or a MDEventWorkspace.
OutputWorkspace(Output:req) Workspace Name of the newly created cloned workspace.
- convertUnits(InputWorkspace, Target, EMode=None, EFixed=None, AlignBins=None, ConvertFromPointData=None)#
Performs a unit change on the X values of a workspace
Property descriptions:
InputWorkspace(Input:req) MatrixWorkspace Name of the input workspace
OutputWorkspace(Output:req) MatrixWorkspace Name of the output workspace, can be the same as the input
Target(Input:req) string The name of the units to convert to (must be one of those registered in the Unit Factory)[DeltaE, DeltaE_inFrequency, DeltaE_inWavenumber, dSpacing, dSpacingPerpendicular, Energy, Energy_inWavenumber, Momentum, MomentumTransfer, QSquared, SpinEchoLength, SpinEchoTime, TOF, Wavelength]
EMode(Input) string The energy mode (default: elastic)[Elastic, Direct, Indirect]
EFixed(Input) number Value of fixed energy in meV : EI (EMode=’Direct’) or EF (EMode=’Indirect’) . Must be set if the target unit requires it (e.g. DeltaE)
AlignBins(Input) boolean If true (default is false), rebins after conversion to ensure that all spectra in the output workspace have identical bin boundaries. This option is not recommended (see http://docs.mantidproject.org/algorithms/ConvertUnits).
ConvertFromPointData(Input) boolean When checked, if the Input Workspace contains Points the algorithm ConvertToHistogram will be run to convert the Points to Bins. The Output Workspace will contains Bins.
- delete(Workspace)#
Removes a workspace from memory.
Property descriptions:
Workspace(Input:req) Workspace Name of the workspace to delete.
- displayNormalization((IMDWorkspace)self) MDNormalization :#
Returns the visual
MDNormalizationof the workspace.
- displayNormalizationHisto((IMDWorkspace)self) MDNormalization :#
For MDEventWorkspaces returns the visual
MDNormalizationof derived MDHistoWorkspaces. For all others returns the same as displayNormalization.
- estimateResolution((MDGeometry)self) numpy.ndarray :#
Returns a numpy array containing the width of the smallest bin in each dimension
- getBasisVector((MDGeometry)self, (int)index) mantid.kernel._kernel.VMD :#
Returns a
VMDobject defining the basis vector for the specified dimension
- getComment((Workspace)self) str :#
Returns the comment field on the workspace
- getDimension((MDGeometry)self, (int)index) mantid.geometry._geometry.IMDDimension :#
Returns the description of the
IMDDimensionat the given index (starts from 0). Raises RuntimeError if index is out of range.
- getDimensionIndexById((MDGeometry)self, (str)id) int :#
Returns the index of the
IMDDimensionwith the given ID. Raises RuntimeError if the name does not exist.
- getDimensionIndexByName((MDGeometry)self, (str)name) int :#
Returns the index of the dimension with the given name. Raises RuntimeError if the name does not exist.
- getDimensionWithId((MDGeometry)self, (str)id) mantid.geometry._geometry.IMDDimension :#
Returns the description of the
IMDDimensionwith the given id string. Raises ValueError if the string is not a known id.
- getGeometryXML((MDGeometry)self) str :#
Returns an XML representation, as a string, of the geometry of the workspace
- getHistory((Workspace)self) WorkspaceHistory :#
Return read-only access to the
WorkspaceHistory
- getMemorySize((Workspace)self) int :#
Returns the memory footprint of the workspace in KB
- getNEvents((IMDWorkspace)self) int :#
Returns the total number of events, contributed to the workspace
- getNPoints((IMDWorkspace)self) int :#
Returns the total number of points within the workspace
- getName((Workspace)self) str :#
Returns the name of the workspace. This could be an empty string
- getNonIntegratedDimensions((MDGeometry)self) list :#
Returns the description objects of the non-integrated dimension as a python list of
IMDDimension.
- getNumDims((MDGeometry)self) int :#
Returns the number of dimensions present
- getNumNonIntegratedDims((MDGeometry)self) int :#
Returns the number of non-integrated dimensions present
- getNumberTransformsFromOriginal((MDGeometry)self) int :#
Returns the number of transformations from original workspace coordinate systems
- getNumberTransformsToOriginal((MDGeometry)self) int :#
Returns the number of transformations to original workspace coordinate systems
- getOrigin((MDGeometry)self) mantid.kernel._kernel.VMD :#
Returns the vector of the origin (in the original workspace) that corresponds to 0,0,0… in this workspace
- getOriginalWorkspace((MDGeometry)self, (int)index) Workspace :#
Returns the source workspace attached at the given index
- getSpecialCoordinateSystem((IMDWorkspace)self) mantid.kernel._kernel.SpecialCoordinateSystem :#
Returns the special coordinate system of the workspace
- getTDimension((MDGeometry)self) mantid.geometry._geometry.IMDDimension :#
Returns the
IMDDimensiondescription mapped to time
- getTitle((Workspace)self) str :#
Returns the title of the workspace
- getXDimension((MDGeometry)self) mantid.geometry._geometry.IMDDimension :#
Returns the
IMDDimensiondescription mapped to X
- getYDimension((MDGeometry)self) mantid.geometry._geometry.IMDDimension :#
Returns the
IMDDimensiondescription mapped to Y
- getZDimension((MDGeometry)self) mantid.geometry._geometry.IMDDimension :#
Returns the
IMDDimensiondescription mapped to Z
- hasOriginalWorkspace((MDGeometry)self, (int)index) bool :#
Returns True if there is a source workspace at the given index
- id((DataItem)self) str :#
The string ID of the class
- isDirty((Workspace)self[, (int)n]) bool :#
True if the workspace has run more than n algorithms (Default=1)
- isGroup((Workspace)self) bool :#
Returns if it is a group workspace
- isMDHistoWorkspace((IMDWorkspace)self) bool :#
Returns True if this is considered to be binned data.
- maskDetectors(Workspace, SpectraList=None, DetectorList=None, WorkspaceIndexList=None, MaskedWorkspace=None, ForceInstrumentMasking=None, StartWorkspaceIndex=None, EndWorkspaceIndex=None, ComponentList=None)#
An algorithm to mask a detector, or set of detectors, as not to be used. The workspace spectra associated with those detectors are zeroed.
Property descriptions:
Workspace(InOut:req) Workspace The name of the input and output workspace on which to perform the algorithm.
SpectraList(Input) int list A list of spectra to mask
DetectorList(Input) int list A list of detector ID’s to mask
WorkspaceIndexList(Input) unsigned int list A list of the workspace indices to mask
MaskedWorkspace(Input) MatrixWorkspace If given but not as a SpecialWorkspace2D, the masking from this workspace will be copied. If given as a SpecialWorkspace2D, the masking is read from its Y values.[]
ForceInstrumentMasking(Input) boolean Works when ‘MaskedWorkspace’ is provided and forces to use spectra-detector mapping even in case when number of spectra in ‘Workspace’ and ‘MaskedWorkspace’ are equal
StartWorkspaceIndex(Input) number If other masks fields are provided, it’s the first index of the target workspace to be allowed to be masked from by these masks, if not, its the first index of the target workspace to mask. Default value is 0 if other masking is present or ignored if not.
EndWorkspaceIndex(Input) number If other masks are provided, it’s the last index of the target workspace allowed to be masked to by these masks, if not, its the last index of the target workspace to mask. Default is number of histograms in target workspace if other masks are present or ignored if not.
ComponentList(Input) str list A list names of components to mask
- name((DataItem)self) str :#
The name of the object
- numOriginalWorkspaces((MDGeometry)self) int :#
Returns the number of source workspaces attached
- readLock((DataItem)self) None :#
Acquires a read lock on the data item.
- setComment((Workspace)self, (str)comment) None :#
Set the comment field of the workspace
- setTitle((Workspace)self, (str)title) None :#
Set the title of the workspace
- threadSafe((DataItem)self) bool :#
Returns true if the object can be accessed safely from multiple threads
- unlock((DataItem)self) None :#
Unlocks a read or write lock on the data item.

