Table of Contents
| Name | Direction | Type | Default | Description |
|---|---|---|---|---|
| Filename | Input | list of str lists | Mandatory | Files to combine in reduction. Allowed extensions: [‘_event.nxs’, ‘.nxs.h5’, ‘.nxs’] |
| FilterByTofMin | Input | number | Optional | Optional: To exclude events that do not fall within a range of times-of-flight. This is the minimum accepted value in microseconds. Keep blank to load all events. |
| FilterByTofMax | Input | number | Optional | Optional: To exclude events that do not fall within a range of times-of-flight. This is the maximum accepted value in microseconds. Keep blank to load all events. |
| FilterByTimeStop | Input | number | Optional | Optional: To only include events before the provided stop time, in seconds (relative to the start of the run). |
| MomentumMin | Input | number | Optional | Minimum value in momentum. This should match the Flux momentum if the output is to be used with MDNorm |
| MomentumMax | Input | number | Optional | Maximum value in momentum. This should match the Fluxmomentum if the output is to be used with MDNorm |
| UBMatrix | Input | string | Path to an ISAW-style UB matrix text file. See LoadIsawUB. Allowed extensions: [‘.mat’, ‘.ub’, ‘.txt’] | |
| SetGoniometer | Input | boolean | False | Set which Goniometer to use. See SetGoniometer |
| Goniometers | Input | string | None, Specify Individually | Set the axes and motor names according to goniometers that we define in the code (Universal defined for SNS). Allowed values: [‘None, Specify Individually’, ‘Universal’] |
| Axis0 | Input | string | Axis0: name, x,y,z, 1/-1 (1 for ccw, -1 for cw rotation). A number of degrees can be used instead of name. Leave blank for no axis | |
| Axis1 | Input | string | Axis1: name, x,y,z, 1/-1 (1 for ccw, -1 for cw rotation). A number of degrees can be used instead of name. Leave blank for no axis | |
| Axis2 | Input | string | Axis2: name, x,y,z, 1/-1 (1 for ccw, -1 for cw rotation). A number of degrees can be used instead of name. Leave blank for no axis | |
| LoadInstrument | Input | string | Load a different instrument IDF onto the data from a file. See LoadInstrument. Allowed extensions: [‘.xml’] | |
| ApplyCalibration | Input | TableWorkspace | Calibration will be applied using this TableWorkspace using ApplyCalibration. | |
| DetCal | Input | string | Load an ISAW DetCal calibration onto the data from a file. See LoadIsawDetCal. Allowed extensions: [‘.detcal’] | |
| CopyInstrumentParameters | Input | MatrixWorkspace | The input workpsace from which CopyInstrumentParameters will copy parameters to data | |
| MaskFile | Input | string | Masking file for masking. Supported file format is XML and ISIS ASCII. See LoadMask. Allowed extensions: [‘.xml’, ‘.msk’] | |
| OutputWorkspace | Output | Workspace | Mandatory | Output Workspace |
| QFrame | Input | string | Q_sample | Selects Q-dimensions of the output workspace. Q (sample frame): Wave-vector converted into the frame of the sample (taking out the goniometer rotation). HKL: Use the sample’s UB matrix to convert Wave-vector to crystal’s HKL indices. Allowed values: [‘Q_sample’, ‘HKL’] |
| Uproj | Input | dbl list | Defines the first projection vector of the target Q coordinate system in Q3D mode - Default (1,0,0) | |
| Vproj | Input | dbl list | Defines the second projection vector of the target Q coordinate system in Q3D mode - Default (0,1,0). | |
| Wproj | Input | dbl list | Defines the third projection vector of the target Q coordinate system in Q3D mode. - Default (0,0,1) | |
| MinValues | Input | dbl list | It has to be N comma separated values, where N is the number of dimensions of the target workspace. Values smaller then specified here will not be added to workspace. Number N is defined by properties 4,6 and 7 and described on MD Transformation factory page. See also ConvertToMDMinMaxLocal v1 | |
| MaxValues | Input | dbl list | A list of the same size and the same units as MinValues list. Values higher or equal to the specified by this list will be ignored | |
| SplitInto | Input | int list | 5 | A comma separated list of into how many sub-grid elements each dimension should split; or just one to split into the same number for all dimensions. Default 5. |
| SplitThreshold | Input | number | 1000 | How many events in a box before it should be split. Default 1000. |
| MaxRecursionDepth | Input | number | 20 | How many levels of box splitting recursion are allowed. The smallest box will have each side length 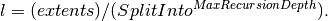 Default 20. Default 20. |
| OverwriteExisting | Input | boolean | True | By default (“1”), existing Output Workspace will be replaced. Select false (“0”) if you want to add new events to the workspace, which already exist. Choosing “0” can be very inefficient for file-based workspaces |
This workflow algorithm loads, converts to MDWorkspace and combines a series of runs. The resulting workspace is a MDWorkspace containing a volume of scattering events. If a UB matrix is provided and QFrame=’HKL’, then the MD workspace will be in HKL otherwise it will be in Q_sample.
This is not a correctly normalized workspace, for that look at SingleCrystalDiffuseReduction. The output workspace can be used with FindPeaksMD and from that determine the UB matrix. The output workspace (if in Q sample) can then be used with MDNorm to correctly bin and normalise the data.
The input filename follows the syntax from MultipleFileProperty
You should refer to ConvertToMD for all its options. If MinValues and MaxValues are not provided then limits will be calculated with ConvertToMDMinMaxGlobal
The resulting workspaces can be saved and loaded with SaveMD and LoadMD respectively.
Single file Q-sample
ConvertMultipleRunsToSingleCrystalMD(Filename='CNCS_7860',
MinValues=[-2,-2,-2],
MaxValues=[2,2,2],
OutputWorkspace='output',
SetGoniometer=True,
Axis0="huber,0,1,0,1")
ws=mtd['output']
print("The workspace is in {}".format(ws.getSpecialCoordinateSystem()))
print("There are {} experiment runs in the workspace".format(ws.getNumExperimentInfo()))
print("Number of Events = {}".format(ws.getNEvents()))
print("There are {} dimensions with names: {} {} {}".format(ws.getNumDims(), ws.getDimension(0).name, ws.getDimension(1).name, ws.getDimension(2).name))
Output:
The workspace is in QSample
There are 1 experiment runs in the workspace
Number of Events = 100210
There are 3 dimensions with names: Q_sample_x Q_sample_y Q_sample_z
Multiple files Q-sample
ConvertMultipleRunsToSingleCrystalMD(Filename='CORELLI_29782:29817:10',
FilterByTofMin=1000,
FilterByTofMax=16666,
OutputWorkspace='output',
SetGoniometer=True,
Axis0="BL9:Mot:Sample:Axis1,0,1,0,1")
ws=mtd['output']
print("The workspace is in {}".format(ws.getSpecialCoordinateSystem()))
print("There are {} experiment runs in the workspace".format(ws.getNumExperimentInfo()))
print("Number of Events = {}".format(ws.getNEvents()))
print("There are {} dimensions with names: {} {} {}".format(ws.getNumDims(), ws.getDimension(0).name, ws.getDimension(1).name, ws.getDimension(2).name))
Output:
The workspace is in QSample
There are 4 experiment runs in the workspace
Number of Events = 47223197
There are 3 dimensions with names: Q_sample_x Q_sample_y Q_sample_z
Single file HKL
# Create a ISAW UB file for the test
import mantid
UBfilename=mantid.config.getString("defaultsave.directory")+"ConvertMultipleRunsToSingleCrystalMDTest.mat"
with open(UBfilename,'w') as f:
f.write("0.0 0.5 0.0 \n")
f.write("0.0 0.0 0.25 \n")
f.write("0.2 0.0 0.0 \n")
f.write("2.0 4.0 5.0 90 90 90 40 \n")
f.write("0.0 0.0 0.0 0 0 0 0 \n")
f.write("\n\nsome text about IPNS convention")
ConvertMultipleRunsToSingleCrystalMD(Filename='CNCS_7860',
MinValues=[-2,-2,-2],
MaxValues=[2,2,2],
OutputWorkspace='output',
SetGoniometer=True,
Axis0="huber,0,1,0,1",
UBMatrix=UBfilename,
QFrame='HKL')
ws=mtd['output']
print("The workspace is in {}".format(ws.getSpecialCoordinateSystem()))
print("There are {} experiment runs in the workspace".format(ws.getNumExperimentInfo()))
print("Number of Events = {}".format(ws.getNEvents()))
print("There are {} dimensions with names: {} {} {}".format(ws.getNumDims(), ws.getDimension(0).name, ws.getDimension(1).name, ws.getDimension(2).name))
Output:
The workspace is in HKL
There are 1 experiment runs in the workspace
Number of Events = 112266
There are 3 dimensions with names: [H,0,0] [0,K,0] [0,0,L]
Multiple files HKL
ConvertMultipleRunsToSingleCrystalMD(Filename='CORELLI_29782:29817:10',
FilterByTofMin=1000,
FilterByTofMax=16666,
OutputWorkspace='output',
SetGoniometer=True,
Axis0="BL9:Mot:Sample:Axis1,0,1,0,1",
UBMatrix="/SNS/CORELLI/IPTS-15526/shared/benzil_Hexagonal.mat",
QFrame='HKL')
ws=mtd['output']
print("The workspace is in {}".format(ws.getSpecialCoordinateSystem()))
print("There are {} experiment runs in the workspace".format(ws.getNumExperimentInfo()))
print("Number of Events = {}".format(ws.getNEvents()))
print("There are {} dimensions with names: {} {} {}".format(ws.getNumDims(), ws.getDimension(0).name, ws.getDimension(1).name, ws.getDimension(2).name))
Output:
The workspace is in HKL
There are 4 experiment runs in the workspace
Number of Events = 47223200
There are 3 dimensions with names: [H,0,0] [0,K,0] [0,0,L]
Python: ConvertMultipleRunsToSingleCrystalMD.py (last modified: 2019-02-05)