IntegratePeaksMDHKL v1#
Summary#
Integrate single-crystal peaks in reciprocal space, for MDHistoWorkspaces.
See Also#
IntegratePeaksHybrid, IntegratePeaksUsingClusters, IntegratePeaksMD, IntegratePeaksCWSD
Properties#
Name |
Direction |
Type |
Default |
Description |
|---|---|---|---|---|
InputWorkspace |
Input |
IMDWorkspace |
Mandatory |
An input Sample MDHistoWorkspace or MDEventWorkspace in HKL. |
DeltaHKL |
Input |
number |
0.5 |
Distance from integer HKL to integrate peak. |
GridPoints |
Input |
number |
201 |
Number of grid points for each dimension of HKL box. |
NeighborPoints |
Input |
number |
10 |
Number of points in 5^3 surrounding points above intensity threshold for point to be part of peak. |
FluxWorkspace |
Input |
An optional input workspace containing momentum dependent flux for normalization. |
||
SolidAngleWorkspace |
Input |
An optional input workspace containing momentum integrated vanadium for normalization (a measure of the solid angle). |
||
PeaksWorkspace |
Input |
PeaksWorkspace |
Mandatory |
A PeaksWorkspace containing the peaks to integrate. |
OutputWorkspace |
Output |
PeaksWorkspace |
Mandatory |
The output PeaksWorkspace will be a copy of the input PeaksWorkspace with the peaks’ integrated intensities. |
BackgroundInnerRadius |
Input |
number |
Optional |
Optional:Inner radius to use to evaluate the background of the peak. If omitted background is region of HKL box - peak. |
BackgroundOuterRadius |
Input |
number |
Optional |
Optional:Outer radius to use to evaluate the background of the peak. The signal density around the peak (BackgroundInnerRadius < r < BackgroundOuterRadius) is used to estimate the background under the peak. If omitted background is region of HKL box - peak. |
Description#
IntegratePeaksMDHKL provides integration of a
MDHistoWorkspace or MDEventWorkspace in 3-dimensions.
The units of the workspace must be HKL. The main usage will be for data normalized by
MDNormSCD.
A 3D box is created for each peak and the background and peak data are separated. The intensity and sigma of the
intensity is found from the grid inside the peak and the background is subtracted. The boxes are created and integrated
in parallel and less memory is required than binning all HKL at once.
\(I_{corr} = I_{peak} - pts_{peak}/pts_{bg} * I_{bg}\)
with the errors summed in quadrature:
\(\sigma_{I,corr}^2 = \sigma_{I,peak}^2 + (pts_{peak}/pts_{bg})^2 * \sigma_{I,bg}^2\)
Using the DeltaHKL parameter, the problem of nearly peaks or regions of diffuse scattering can be avoided. Also for normalized data, the unmeasured data points are excluded from the background. See white regions in last figure.
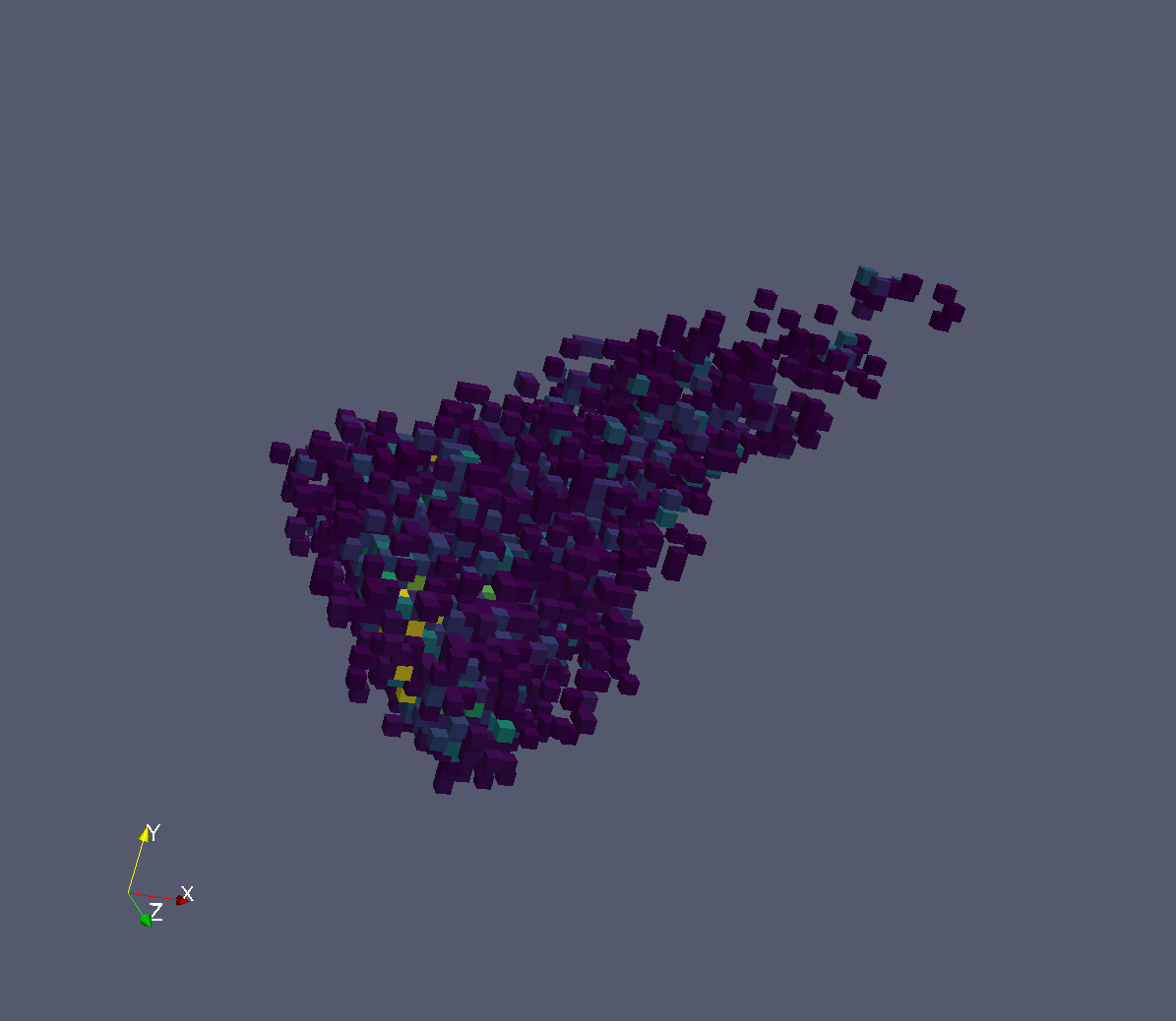
Peak Integration Input. 3D Box.#
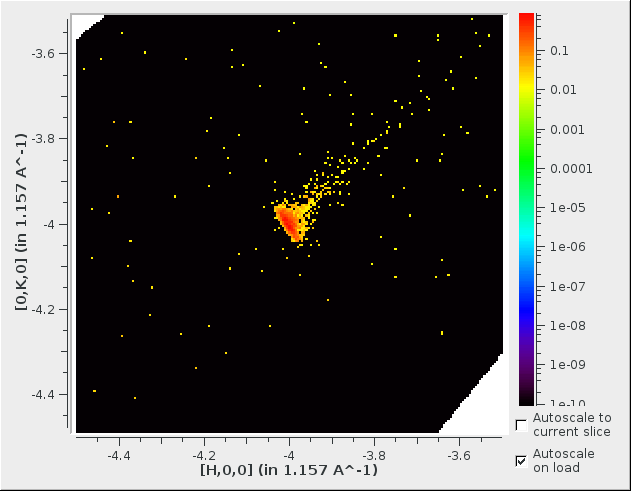
Integration slice at center of box.#

Integration slice of peak grid points.#
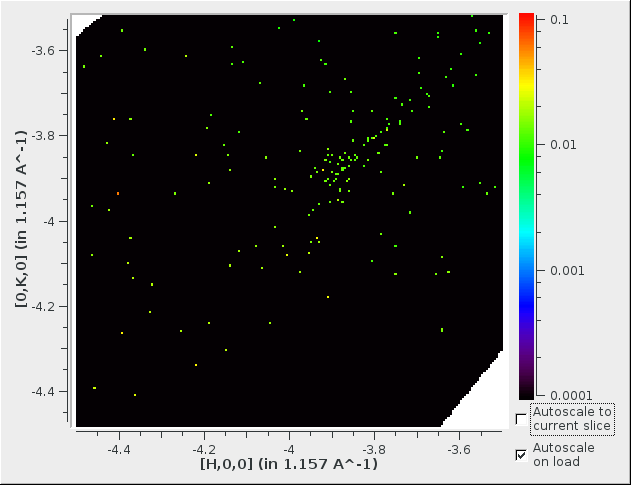
Integration slice of background grid points.#
Usage#
Example - IntegratePeaksMDHKL event histo
#Create PeaksWorkspace
sampleWs = CreateSampleWorkspace()
pws = CreatePeaksWorkspace(InstrumentWorkspace=sampleWs,NumberOfPeaks=3)
p = pws.getPeak(0)
p.setHKL(5,0,0)
p = pws.getPeak(1)
p.setHKL(0,0,0)
p = pws.getPeak(2)
p.setHKL(-5,0,0)
#Test with MDEventWorkspace
mdws = CreateMDWorkspace(Dimensions=3, Extents=[-10,10,-10,10,-10,10], Names='[H,0,0],[0,K,0],[0,0,L]',Units='A^-1,A^-1,A^-1',Frames='HKL,HKL,HKL')
FakeMDEventData(InputWorkspace=mdws, PeakParams=[100000,-5,0,0,1])
FakeMDEventData(InputWorkspace=mdws, PeakParams=[100000,0,0,0,1])
FakeMDEventData(InputWorkspace=mdws, PeakParams=[100000,5,0,0,1])
pws =IntegratePeaksMDHKL(InputWorkspace=mdws,PeaksWorkspace=pws,DeltaHKL=1.5,GridPoints=21)
for i in range(3):
p = pws.getPeak(i)
print('{:.7f} {:.9f}'.format(p.getIntensity(),p.getSigmaIntensity()))
#Test with MDHistoWorkspace
mdws = BinMD(InputWorkspace=mdws,AlignedDim0="[H,0,0],-10,10,101",AlignedDim1="[0,K,0],-10,10,101",AlignedDim2="[0,0,L],-10,10,101")
pws =IntegratePeaksMDHKL(InputWorkspace=mdws,PeaksWorkspace=pws,DeltaHKL=1.5,GridPoints=21)
for i in range(3):
p = pws.getPeak(i)
print('{:.7f} {:.9f}'.format(p.getIntensity(),p.getSigmaIntensity()))
Output:
99965.2588423 316.186057523
99965.2588423 316.186057523
99965.2588423 316.186057523
99959.4727293 316.176064633
99978.7624499 316.200132638
99961.9286227 316.179180305
Categories: AlgorithmIndex | MDAlgorithms\Peaks | Crystal\Integration
Source#
C++ header: IntegratePeaksMDHKL.h
C++ source: IntegratePeaksMDHKL.cpp

